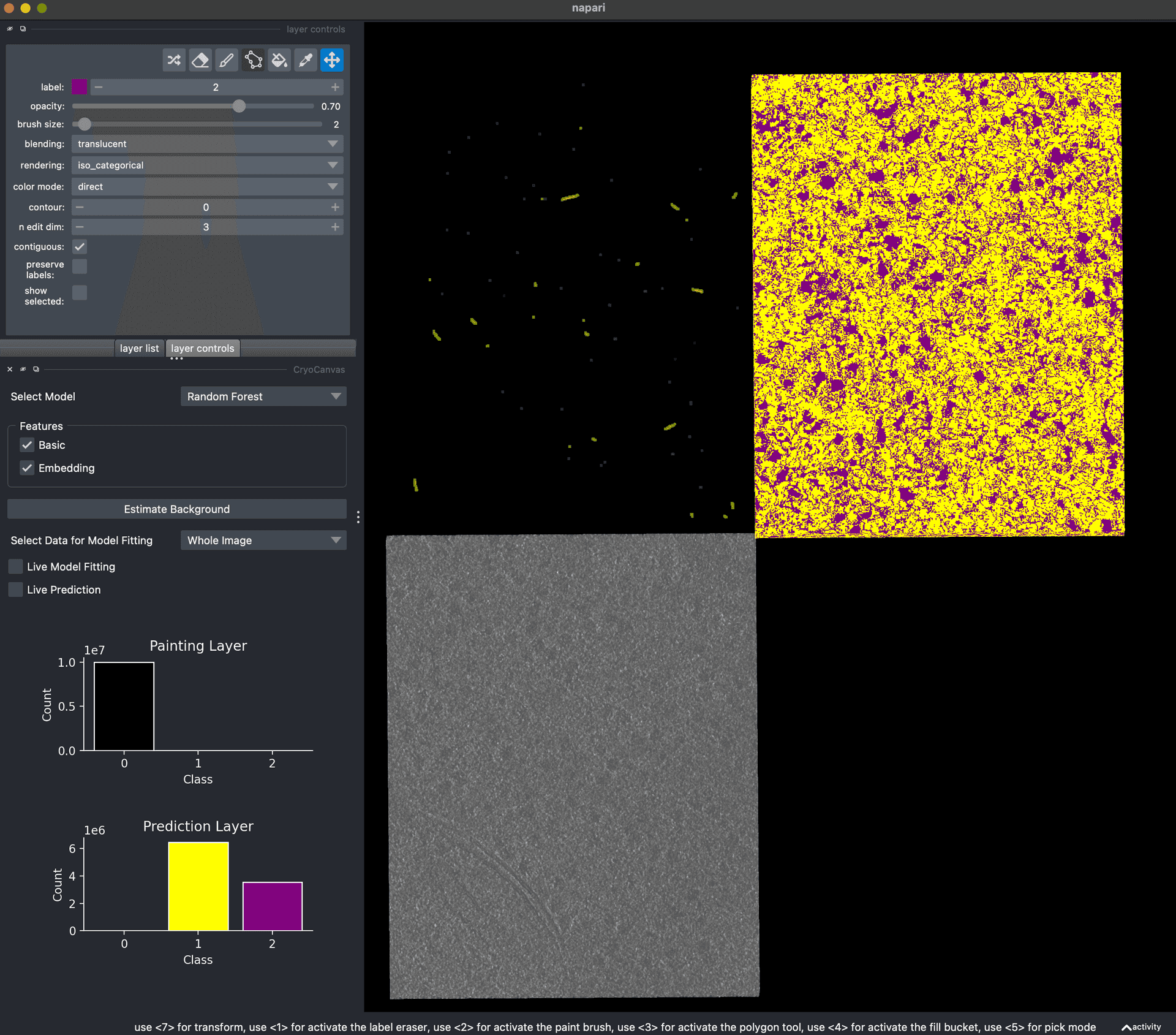
- First portable CellCanvas demo.First portable CellCanvas demoKyle Harrington.
![CelCanvas screenshot.]()
- Parent environment for supporting the Winter 2024 UX Evaluation.Parent environment for supporting the Winter 2024 UX EvaluationKyle. Also check out TeamTomo
- Fetch datasets for the UX evaluationFetch datasets for the UX evaluation in Winter 2024.
- Run UX Evaluation.Run the UX evaluation itselfKyle Harrington.
- Export results of the UX evaluationExport results of the UX evaluation.
- Predict a Multilabel Segmentation Using a ModelA solution that predicts segmentation using a model for a Copick project and saves it as 'predictionsegmentation'.
- Run CellCanvas.Run CellCanvasCellCanvas team.
- Run SurforamaRun Surforama.CellCanvas team.
- Predict Tomogram Embeddings with SwinUNETR using Copick APIApply a SwinUNETR model to a tomogram fetched using the Copick API to produce embeddings, and save them in a Zarr.
- Batch Process Zarr Files for Pixel EmbeddingAutomatically process all Zarr files within a specified directory structure using a SwinUNETR model.CellCanvas and Copick teams.
- Generate a tomogram with polnetGenerate tomograms with polnet, and save them in a Zarr.Martinez-Sanchez A.*, Jasnin M., Phelippeau H. and Lamm L. Simulating the cellular context in synthetic datasets for cryo-electron tomography, bioRxiv.
- Fetch Data from CryoET Data Portal and Integrate with CopickFetches tomograms, tilt series, annotations, segmentations, and metadata from cryoet_data_portal and integrates them into the specified Copick project.Cellcanvas and Copick teams
- Convert a mrc to omezarr using mrc2omezarrConvert a mrc to omezarr using mrc2omezarr.Utz Ermel.
- Run CellCanvas with a copick project.Run CellCanvas with a copick projectCellCanvas team.
- Run CellCanvas with a copick project.Run CellCanvas with a copick projectCellCanvas team.
- Run CellCanvas with a copick project.Run CellCanvas with a copick projectCellCanvas team.
- Analyze Picks and Corresponding Embeddings for a Single RunGenerates a DataFrame from picks and their corresponding embeddings for a single run and saves it.
- Classify and Match Embeddings to Known Particle Classes with Multithreading Across Multiple DirectoriesUses multithreading to compare median embeddings from multiple Zarr datasets to known class medians and identifies matches based on a configurable distance threshold.
- Paint Copick Picks into a Segmentation LayerA solution that paints picks from a Copick project into a segmentation layer in Zarr.
- Train Random Forest on Zarr Data with Cross-ValidationA solution that trains a Random Forest model using data from a Zarr zip store, filters runs with only one label, and performs cross-validation.
- Generate Multiscale Basic Features with Scikit-Image using Copick API (Chunked, Corrected)Compute multiscale basic features of a tomogram from a Copick run in chunks and save them using Copick's API.
- Process Copick Runs and Save Features and LabelsA solution that processes all Copick runs and saves the resulting features and labels into a Zarr zip store.
- Optimize Random Forest with Optuna on Zarr DataA solution that optimizes a Random Forest model using Optuna, data from a Zarr zip store, and performs 10-fold cross-validation.
- Predict Binary Segmentations Using ModelsA solution that predicts binary segmentations for each label using models created by an optimization solution, and saves them separately.
- Extract Centroids from Multilabel SegmentationA solution that extracts centroids from a multilabel segmentation using Copick and saves them as candidate picks.
- Train LightGBM on Zarr Data with Cross-ValidationA solution that trains a LightGBM model using data from a Zarr zip store, filters runs with only one label, and performs 10-fold cross-validation.
- Train XGBoost on Zarr Data with Cross-ValidationA solution that trains an XGBoost model using data from a Zarr zip store, filters runs with only one label, and performs 10-fold cross-validation.
- Optimize XGBoost with Optuna on Zarr DataA solution that optimizes an XGBoost model using Optuna, data from a Zarr zip store, and performs 10-fold cross-validation.
- Predict a Multilabel Segmentation Using a ModelA solution that predicts segmentation using a model for a Copick project and saves it as 'predictionsegmentation'.
- Compare Picks from Different Users and Sessions with F-beta ScoreA solution that compares the picks from a reference user and session to a candidate user and session for all particle types, providing metrics like average distance, precision, recall, and F-beta score. Computes micro-averaged F-beta score across all runs if run_name is not provided.
- Analyze Median Embeddings for Each Object Type Across Multiple RunsGenerates a file containing the median embeddings for each object type based on the picks in multiple runs, filtered by user IDs if provided.
- Create Picks with Distance to Median EmbeddingCreates a new set of picks for a new session ID, containing the same locations but including the distance to the median embedding in the 'score' attribute.
- Generate a PNS density image from PDB IDGenerate a Point Normal Surface (PNS) file from a PDB ID.Cellcanvas and copick teams
- Evaluate Picks Against Multilabel SegmentationA solution that evaluates picks from a Copick project against a multilabel segmentation and computes metrics for each (user_id, session_id, object_name) pair for each run and across all runs.
- Split Dataset for Training and TestingA solution that splits datasets into training and test sets, ensuring distributions are preserved.
- Train SwinUnetr Pixel Embedding NetworkTrain the SwinUnetr pixel embedding network using the provided script and dataset.Morphospaces team.
- Run CoPick Live.Run CoPick LiveCoPick Live team.
- Grid Picks from TomogramA solution that places a grid of picks based on the radius of each pickable object, parameterized by a multiple of the particle radius, using tomogram shape.
- Submit Album Job ArraySubmit another album solution to Slurm as a job array by using the runs in a Copick project.
- Train UNet Model using MONAI with Multiple Runs and MLflowTrain a UNet model to predict segmentation masks using MONAI from multiple runs with MLflow tracking.
- Display Copick Project IndexA solution that opens a Copick project and displays the index using textual.
- Voxel Counts per LabelA solution that counts the number of voxels per label in a segmentation and saves the results as a CSV and HTML page.
- Napari Copick Plugin LauncherA solution that installs napari-copick and launches the CopickPlugin.
- Compare All Picks from Different Users and SessionsA solution that uses the compare-picks album solution to evaluate all user_id and session_id pairs listed in a CSV file, creating JSON output files for each pair in a specified directory and submitting jobs to Slurm.
- Train SwinUnetr Pixel Embedding Network with Mrc DtataloaderTrain the SwinUnetr pixel embedding network using the provided script and dataset.Morphospaces team.
- Compare Rankings from Different RunsA solution that compares the rankings of candidates in the public and private test sets using various rank metrics.
- Train SwinUNETR Pixel Embedding Network with Copick DatasetTrain the SwinUNETR pixel embedding network using the Copick dataset.Morphospaces team.
- Train 3D UNet for Segmentation with Copick DatasetTrain a 3D UNet network using the Copick dataset for segmentation.Morphospaces team.
- Export Copick Runs to HDF5A solution that exports multiple Copick runs' tomograms and picks into an HDF5 file.
- Train 3D UNet for Segmentation with Copick DatasetTrain a 3D UNet network using the Copick dataset for segmentation.Morphospaces team.
- Train 3D ResUNet for Segmentation with Copick DatasetTrain a 3D ResUNet network using the Copick dataset for segmentation.Morphospaces team.
- Train 3D Swin UNETR for Segmentation with Copick DatasetTrain a 3D Swin UNETR network using the Copick dataset for segmentation.Morphospaces team.
- An environment to support copick monai projectsAn album solution for copick monai morphospaces projects .Kyle Harrington
- Generate Segmentation Masks using UNet CheckpointGenerate segmentation masks using a trained UNet checkpoint on the Copick dataset.Cellcanvas team.
- An environment to support polnet and copickAn album solution for copick polnet projects .Kyle Harrington
- Generate segmentation masks for Copick Dataset using Segment Anything 2Automatically generate segmentation masks for the Copick dataset. Install on a device with GPU access. Need cuda/12.1.1_530.30.02 and cudnn/8.9.7.29_cuda12 to compile.Meta AI team.
- Generate a Tensorboard projector for visualzing the embeddingsAutomatically generate Tensorboard event files and launch a visualizer for it. Currently suppot zarr and npy files.Tensorboard Team.
- FastAPI Album ServerA FastAPI server to manage Album solutions.
- FastAPI CellCanvas ServerBackend for CellCanvas with Copick Config Support.
- napari-CellCanvasA solution that launches napari-cellcanvas with optional config fetching.
- Fetch Copick Config and Write to FileFetches a Copick config from a FastAPI server and writes it to a file.
- Display Copick Project IndexA solution that opens a Copick project and displays the index using Panel.
- Train Binary SwinUNETR Pixel Embedding Network with Copick DatasetTrain the Binary SwinUNETR pixel embedding network using the Copick dataset.Pixeltapestry team.
- Generate a PNS density image from PDB IDGenerate a Point Normal Surface (PNS) file from a PDB ID.Cellcanvas and copick teams
- Generate Multiscale Basic Features with Torch using Copick API (Chunked, Corrected)Compute multiscale basic features of a tomogram from a Copick run in chunks and save them using Copick's API.
- Mock Annotation and XGBoost Training on Copick DataA solution that creates mock annotations based on multilabel segmentation, trains XGBoost models in steps, and generates predictions.
- Model Evaluation on Copick DataEvaluates segmentation models from the mock-annotation solution on Copick data, generating metrics like IoU and F1, and saves the predicted segmentation into the Copick project.
- Mock Annotation and PyTorch Training on Copick DataA solution that creates mock annotations based on multilabel segmentation, trains a PyTorch segmentation model in steps, and generates predictions.
- FastAPI CellCanvas ServerBackend for CellCanvas with Copick Config Support.
- Train XGBoost on Copick Data with Cross-ValidationA solution that processes Copick runs, filters runs with only one label, and trains an XGBoost model with 10-fold cross-validation.